COVID-19 drug targets: understanding endocytosis
With regards to the COVID-19 pandemic, several drugs and vaccines are still in the pre-clinical setting and very few have received approval from the Food and Drug Administration (FDA), or equivalent international regulatory bodies, to be tested in human clinical trials (more info at https://clinicaltrials.gov/). In a recent Viewpoint article in The FEBS Journal, Oleg Glebov (King’s College London, UK, and Qingdao University Medical School, China) underscores that despite 71 potential vaccines under development, only five are at human clinical trial stage [1]. Therefore currently, palliative care remains the only option to patients. To expedite the typical 15-year track of a therapeutic from discovery through to commercial availability (timeline in Fig. 1), a COVID-19 accelerated drug development initiative has been announced to shorten this to 18 months.
.png)
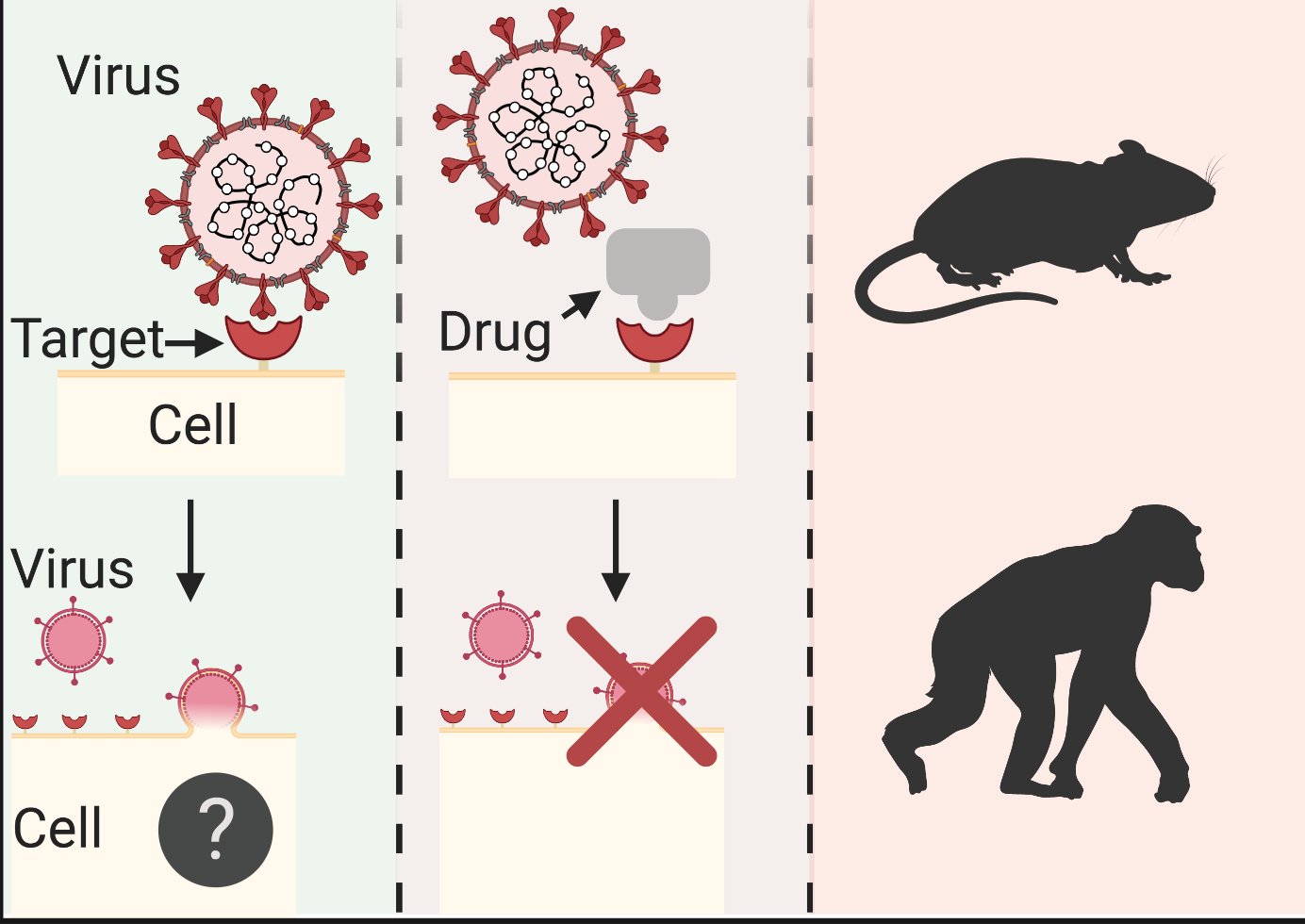
Figure 1. Typical route for rational drug discovery. (A) Once a firm grasp of the biological process under study is attained, an array of candidate drugs can be rationally designed to interfere with said process. The safety and efficacy of the most promising candidates would be initially examined in various animal models; (B) to ensure these findings hold in humans, a few lead candidates would be trialled through three distinct phases, addressing toxicity, potency and reproducibility; (C) fulfilling all previous criteria, an expert regulatory body scrutinises clinical trial data and decides on approving a promising therapeutic. Question mark indicates exact mechanism of biological process is not yet understood. Grey puzzle piece denotes a potential therapeutic agent blocking access to a cell surface protein (red crescent) from a coronavirus. Coloured human population cartoon distinguishes healthy (teal) from infected (red) individuals. Adapted from https://www.nebiolab.com/drug-discovery-and-development-process/, created with BioRender.
Where do we turn our attention for drug discovery?
In the debate of whether to target the human body or part of the novel coronavirus for drug development, Glebov cites the high mutation rate of the virus as a drawback against the latter. Instead, drug resistance is less likely to arise by targeting the human body, treatments identified could be applicable to other viral infections if a similar mechanism is used and it could make pre-clinical drug screening easier due to the use of a single cell model. Namely, the pre-clinical set up would include a cell model of the host grown in a dish, into which various drugs would be introduced, following the addition of a virus or viral components. Interfering with physiological processes of the host cell (humans) is not without shortcomings, however, as it could result in undesirable side-effects due to the drug(s) affecting other biological processes that might be linked.
Drugging endocytosis: targeting the vehicle of the virus in transit
Analogous to a key-and-lock scenario, upon viral attachment to a host cell surface via specific spike proteins on the viral envelope, the virus can be licensed for internalisation through endocytosis (Fig. 1A). However, the result of the endocytic process will depend on the type of outer cell surface protein bound by the virus. The exact internalisation mechanism hijacked by the virus still remains elusive. Data suggest that even though infection might start at nasal epithelial cells, once the virus reaches the lower airways of the lung, a surge of cytokines, molecules associated with inflammation, escalates infection symptoms. Hence, it remains to be seen whether intercepting viral entry at those locations would be clinically beneficial.
How can we experimentally dissect the intracellular viral journey?
In his Viewpoint article, Glebov lists various experimental approaches to address key questions regarding viral endocytosis such as:
1. time taken from viral attachment onto cell surface to cell cytosol arrival
2. mechanism of transport, and
3. distinguishing if viral transport competes with simultaneous transport of necessary molecules to the cell.
These questions are particularly relevant to the choice of cell model for such investigations as well as drug screening. For instance, nasal epithelial cells would have a distinct abundance of endocytosis-mediating proteins compared to cells deeper in the lung airway e.g. pneumocytes. This means that not all endocytic pathways could be modelled in all cell types.
An option for fast-track drug discovery?
Glebov proposes a list of 11 drugs (previously approved by the FDA for other diseases) to be tested for their endocytosis-blocking potential. Repurposing such available medicine could ‘fast-track’ drug discovery, since some have exhibited obstruction of endocytic pathways in cell models. Again, given the unclear mechanism behind SARS-CoV-2 endocytic mechanism(s), more testing might be needed. Overall, Glebov brings together several studies to make suggestions of tools and models available to consolidate SARS-CoV-2 basic biology. In parallel, the combination of drug design endeavours and existing drug repurposing should push our understanding of endocytosis exploited by the virus farther and at an accelerated rate, hopefully also guiding drug development.
The Viewpoint article can now be read in full in The FEBS Journal: https://febs.onlinelibrary.wiley.com/doi/full/10.1111/febs.15369
References:
1. Glebov, O.O. (2020), Understanding SARS‐CoV‐2 endocytosis for COVID‐19 drug repurposing. FEBS J. doi:10.1111/febs.15369




Join the FEBS Network today
Joining the FEBS Network’s molecular life sciences community enables you to access special content on the site, present your profile, 'follow' contributors, 'comment' on and 'like' content, post your own content, and set up a tailored email digest for updates.