How can artificial intelligence contribute to explaining biological mechanisms by which an environmental chemical may cause health disorders?

The Exposome concept
Every day, all of us breathe, eat and drink, as well as carry out a range of unconscious vital functions, such as beating our heart. What many of us don't realize is that with every breath of air, sip of water or nutrient ingested, there is a probability of being exposed to hazardous substances, e.g., pesticides from air or food contaminants such as endocrine disruptors. The “genome” term is well known to the general population, who understand it as all the genes that make us up. However, not all diseases are linked to the genome and some of them need to be triggered by external factors. In 2005, Christopher Wild introduced the “exposome” notion (1), which describes all the environmental factors (i.e., non-genetic) we face from birth to death. The exposome includes chemical, physical, biological, and social stresses (see Figure 1), and contains various sorts of exposure, from air pollution to mental load and sleep quality, through exposure to the sun (ultraviolet lights).
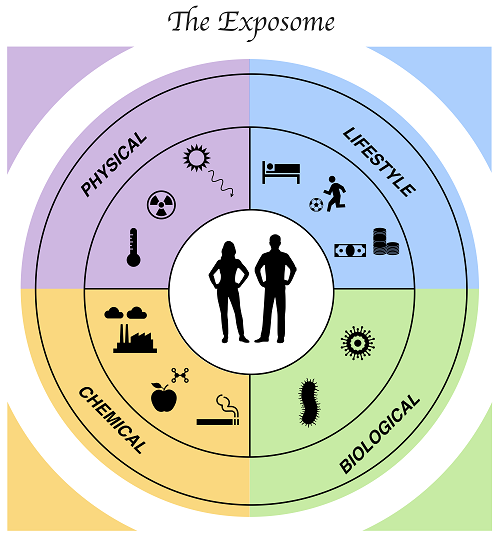
The Adverse Outcome Pathway (AOP) concept
To understand and model how these stresses may cause diseases, the Adverse Outcome Pathway (AOP) concept was formalized in 2010 by G.T. Ankley and his colleagues (2). An AOP is a comprehensive framework that allows the description of a toxicity pathway from a Molecular Initiating Event (MIE) to an Adverse Outcome (AO), passing through some Key Events (KE). Although the stressors that trigger the MIE do not belong to the AOP concept, a MIE must always arise from a stress exposure (chemical, biological, physical, social). Τhe AO can occur at several levels of organization, such as the individual, the population and even the ecosystem (see Figure 2).
%20framework.jpg)
Actually, an AOP can illustrate how a mundane microscopic exposure (e.g., eating an apple with a few pesticides or passive smoking might seem insignificant) may lead to a major macroscopic impact. It is a bit like trying to understand how we missed our flight despite only waking up 5 minutes late. An initial delay of 5 minutes may seem insignificant, but it can lead us to being in a hurry and forgetting our passport, having to turn back to get it, and therefore being not 5 minutes but 10 minutes late. Then, being stuck in traffic jams because of an incident that has just occurred and missing our train and having to take the next one, which means we are now 30 minutes late, and so on. Such an analogy helps to explain how an initial delay of 5 minutes can lead to missing a flight while we had made sure to arrive 2 hours in advance. It is important to note that an AOP is not deterministic, meaning that it is not because you are exposed to the initial stress that you will trigger the pathology, just as it is not because you wake up 5 minutes late that you will necessarily miss your flight. In this example, the MIE is the wake-up delay and the reason for it, whatever it may be, is not part of the AOP.
The AOP-helpFinder tool
The principle of AOP is to collect as much data as possible in order to build up the most realistic model possible. To deal with this issue, the AOP-helpFinder tool is an artificial intelligence-based algorithm (text-mining) used to support the development of AOPs. Nowadays, a huge amount of biological data is gathered in a database of published scientific information, called PubMed, that contains more than 35 million articles. The AOP-helpFinder tool screens automatically all the available literature from this database to find links between stressors and events, and between two biological events.
Briefly, each abstract is first simplified to make it machine-readable by removing among others the stop-words (linking words not required to understand a sentence, e.g., a, the, and, etc.) and then performing a lemmatization or stemming process to simplify words by taking their base or root forms. For example, ‘simple’ or ‘simpl’ are derived from simplify or simplification; another example is the word “leaves”: the base form (lemmatization) is “leaf” while the root form (stemming) is “leav” which may be confused with the verb “to leave”. This second example shows why the lemmatization process is the most powerful.
Once this step has been completed, AOP-helpFinder searches within each abstract for the words of interest (biological events) and then computes scores based on graph theory, as the processed abstract can be considered as an acyclic graph. On the one hand, it focuses on the word position to avoid returning links at the head of the text that may refer to the working hypothesis rather than results (e.g., at the beginning of the abstract, the sentences look something like “The association of BPA and phthalates with breast cancer remains conflicting. This study aims to investigate...” while at the end they resemble “Each 1-unit increase in log-transformed urinary BPA was associated with a 54 % increased breast cancer risk”).
On the other hand, based on Dijkstra’s algorithm (a method to identify the shortest path between nodes in a network, where nodes are words in the case of AOP-helpFinder) it computes a score between words to determine whether the two terms have a reasonable probability of being biologically related (e.g., breast and cancer). If these two aspects are good enough, then the link is considered plausible and included in the results, otherwise the link is not retained. Then, regardless of the success of the first abstract, another one is assessed using the same method, and so on. Finally, once AOP-helpFinder has finished scanning all the abstracts of interest, it computes a confidence score based on a Fisher’s exact test (see Figure 3 for a brief definition) to weight each association (the Key Event Relationships) in order to support the weight of evidence which is an essential feature of AOPs (see Figure 3).
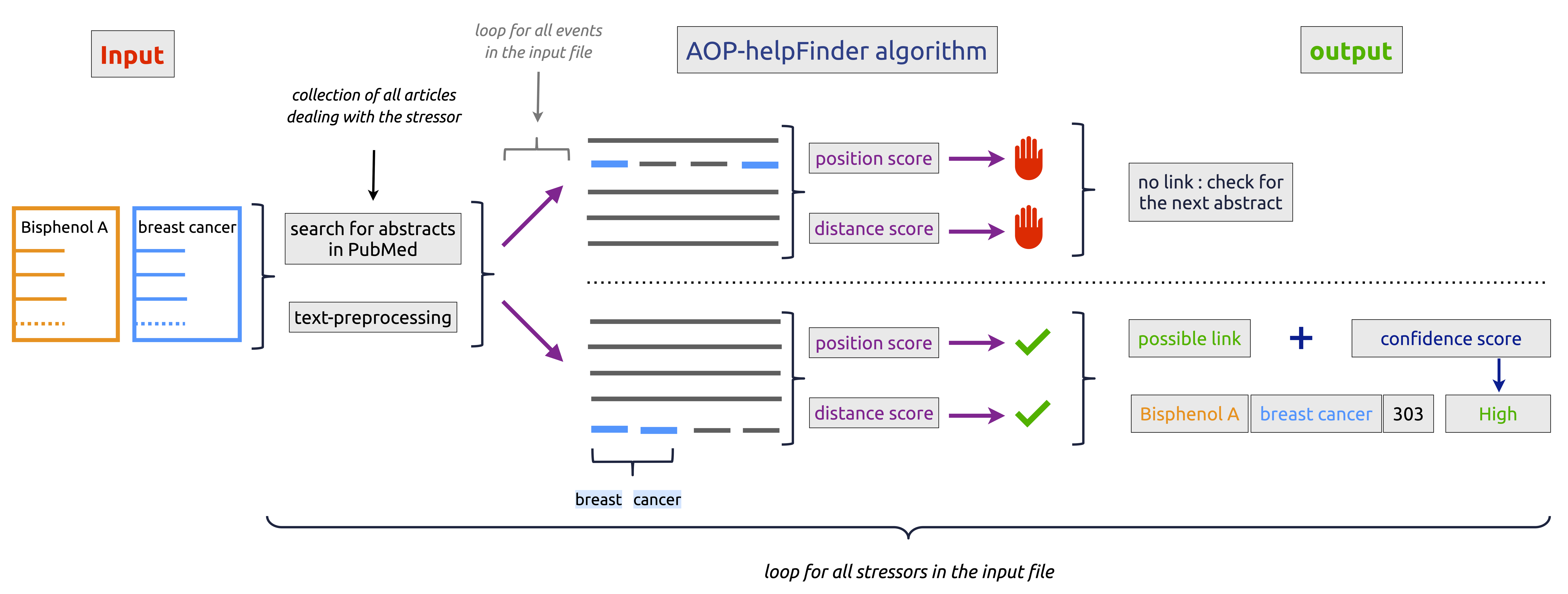
By leveraging a comprehensive analysis of scientific literature, AOP-helpFinder has successfully identified 303 relevant articles with a high confidence score, investigating the association between Bisphenol A, a synthetic chemical commonly found in both food and non-food plastics, and breast cancer (see Figure 3). AOP-helpFinder has also contributed significantly to the development of AOPs in various fields, including the investigation of neurodevelopmental effects resulting from exposure to ionizing radiation (AOP 441) or different types of agrochemicals (AOP 490). Furthermore, this tool has been employed to understand the mechanisms linking dioxins – chemicals generated by combustion or pyrolysis (waste incinerators or forest fires) commonly found in both meat and fish – to breast cancer (AOP 439). These findings demonstrate the utility of AOP-helpFinder in facilitating the exploration of interconnected pathways and providing valuable insights into the relationships between specific chemical exposures and adverse health outcomes. For all those interested in the development of AOPs, the AOP-helpFinder tool is freely available online at the following address: https://aop-helpfinder.u-paris-sciences.fr/
The ongoing advancements in computing power and the abundance of available data have propelled the progress of algorithms in bioinformatic, allowing them to achieve unprecedented levels of performance, in addition to providing ethical alternative methods to animal testing. The more data these in silico models are supplied with, the more accurate they can be expected to be. Besides, as the exposome is made up of a large number of variables, such models are the key to understanding the effect of mixtures on our health by taking into account all the types of stress to which we are subjected.
References
- Wild C. P. (2005). Complementing the genome with an "exposome": the outstanding challenge of environmental exposure measurement in molecular epidemiology. Cancer epidemiology, biomarkers & prevention : a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology, 14(8), 1847–1850. https://doi.org/10.1158/1055-9965.EPI-05-0456
- Ankley, G. T., Bennett, R. S., Erickson, R. J., Hoff, D. J., Hornung, M. W., Johnson, R. D., Mount, D. R., Nichols, J. W., Russom, C. L., Schmieder, P. K., Serrrano, J. A., Tietge, J. E., & Villeneuve, D. L. (2010). Adverse outcome pathways: a conceptual framework to support ecotoxicology research and risk assessment. Environmental toxicology and chemistry, 29(3), 730–741. https://doi.org/10.1002/etc.34
Photo de Luc Tribolet sur Unsplash





Join the FEBS Network today
Joining the FEBS Network’s molecular life sciences community enables you to access special content on the site, present your profile, 'follow' contributors, 'comment' on and 'like' content, post your own content, and set up a tailored email digest for updates.