The Personal Health Train - Secure analysis of large-scale, distributed biomedical data

The importance of large-scale data analysis, also in the context of health data and in the framework of public health institutions, has become abundantly clear during the last year. However, data protection concerns often make a centralized data analysis platform unfeasible. Therefore, the German national medical informatics initiative (MII) chose to prioritize the development of distributed learning frameworks for data analysis. As part of this initiative the Personal Health Train (PHT), a tool for distributed analysis, has been developed.
To streamline development, a FAIR Implementation Network (IN) was founded, which in its manifesto includes the principles guiding the PHT development. The German chapter of the PHT Implementation Network – including the Universities of Aachen, Mittweida, and Tübingen – collaborates to further the development of the PHT and works on case examples, which showcase the functionality of the tool.
What is the core idea of the PHT?
The basic idea of the PHT is that the analysis algorithms (wrapped in a ‘train’) travel between multiple hospitals (so-called ‘train-stations’), which host the input data and computation resources inside their own secure facilities. Trains are implemented as light-weight containers enabling the rapid deployment of complex software pipelines by removing the need to install additional software that might be required to perform the analysis. Such pipelines can range from heavy workloads such as image analysis to bioinformatics pipelines (e.g., nf-core) on high-volume data to secure count queries or basic statistics over multiple sites. Trains travel iteratively from one station to another updating the results securely, which is also known as online learning.
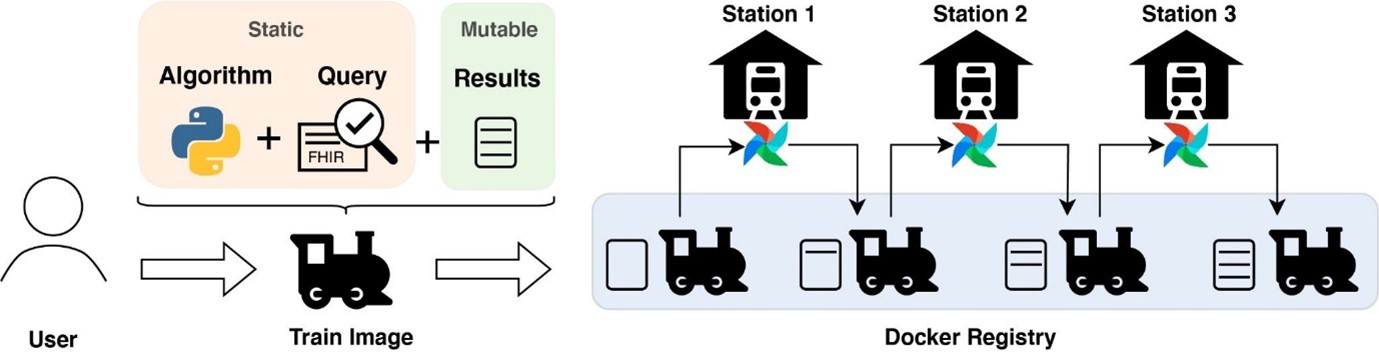
To initiate an analysis the user specifies a search query within the User Interface. To guarantee that data is provided in the correct format the PHT makes use of the FHIR standard. The search query defines which data should be accessed at each location. But trains are capable of processing arbitrary input data. Furthermore, the user uploads the analysis algorithm to be executed at each site. During submission, the signing of hashes with the users' private key guarantees that the submitted files are not modified. These files are added to the train image, which includes the software requirements, dependencies and operation logic. Algorithms and queries are static, while results update iteratively during each execution at the stations. Stations are based on Apache Airflow allowing persistent execution of trains. Within airflow, workflows are defined as Directed Acyclic Graphs (DAG), collecting and running tasks, organized with dependencies.
Different implementations on an compatibility drive
Currently, several implementations exist within the FAIR IN, with the aim to reach compatibility within the next months. Below we detail the status and focus of the Tübingen PHT implementation.
The key focus of the PHT Tübingen is on security. This solution implements security by design. A Security Protocol secures the code executed at each train station against manipulation at any stage. Furthermore, results are encrypted to protect from interception. Additionally, only the analyst and participating study sites can decrypt the results, adding additional security. Secured privacy-preserving count queries are an additional feature that show the possibility of how the PHT architecture can be extended with data privacy-preserving methods (e.g., Paillier cryptosystem).
The PHT Tübingen is deployment-ready and can be operated with patient data. The architecture is open-source, available on Github and consists of several services.
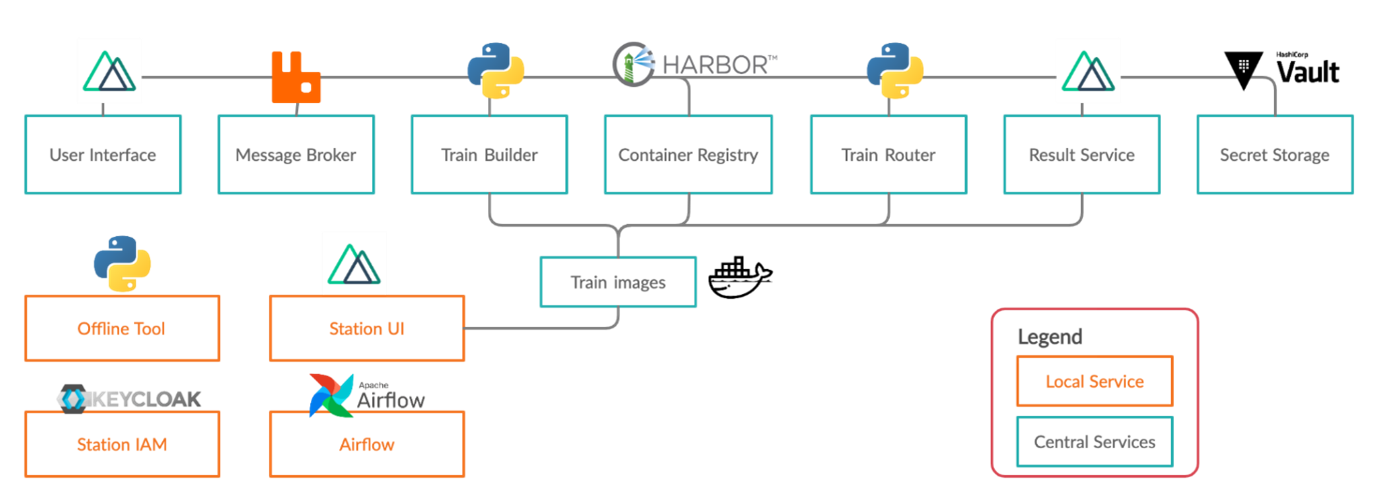
Central service components are used for train submission and train distribution, while local services at hospitals permit the secure execution of trains and reliable data access. Several well-established open-source services are combined with self-developed services. The icons of the services represent the programming language or the software used. Additional information about Tübingen's development can be found on our homepage.
PADME PHT follows the same main principles. The key focus of the PADME PHT ecosystem – developed by Fraunhofer FIT, RWTH Aachen, Cologne, and Leipzig – is on transparency and interoperability of Distributed Analytics platforms. In order to achieve this, “DAMS: A Distributed Analytics Metadata Schema”, which provides a vocabulary to enrich the involved entities with descriptive semantics, is defined and applied. PADME offers additional features like standalone station registry, train execution monitoring tool and visualisation of the updated train contents. These foundations and features are enablers for interoperability while being compliant with FAIR principles.
Future steps and vision
In the following months, both PHT implementations will be used in several projects (CORD, POLAR, Leuko-Expert) on a German national level. These projects will showcase the usability, functionality and interoperability of the PHT infrastructure. The aim is to further extend compatibility to PHT implementations from other countries as well (e.g., Dutch PHT).
Besides, each PHT implementation will focus on additional projects: with Tübingen prioritizing clinical use cases related to Parkinson Disease and Multiple Sclerosis, with a strong focus on imaging data, and PADME contributing to NFDIA4Health for federated learning and FAIR Data Spaces to analyse research and industrial data served in GAIA-X and NFDI.
Another crucial step will be the extension of PHT functionalities: the next major release from Tübingen will extend the current PHT implementation to enable parallel execution of trains and training of global models within a federated learning PHT.
However, the current version is excellent for heavy-work pipelines on high-volume data (omics, raw lab results, different file formats, etc.). Data can remain under the control of sites or laboratories that curated it, guaranteeing only analyses following the protocol can be performed. Especially in extensive, multicentric studies with different stakeholders, data providers lose control as soon data is sent to a single analysis site. Also, if the consent to use the data is withdrawn, any site can remove the subject from its data storage and guarantee no further analysis will be made, including this data.
PADME PHT is being extended to provide a solution for federated learning that enables multiple parties collaboratively training machine learning models. In this solution, the PADME central server orchestrates the federated learning process, including client selection, model broadcasting, client model training, model aggregation and model update. Models instead of raw data are shared during the training process. Differential privacy is applied to provide a high level of robustness and privacy guarantees.
Besides deploying the current architecture to operate on patient data in many German university hospitals, we want to grow a big open-source user community. Only with brilliant ideas and input on desired functionalities can great software be developed. Please contact us if you are interested in joining the PHT community and stay in touch for future updates.
This post was written by Marius de Arruda Botelho Herr, at DIFUTURE Tübingen, University Hospital Tübingen, Medical Data Integration Center, meDIC, with valuable contributions from Stephanie Biergans, Michael Graf, Yeliz Ucer, and Oya Beyan.
Photo by Mathew Schwartz on Unsplash





Join the FEBS Network today
Joining the FEBS Network’s molecular life sciences community enables you to access special content on the site, present your profile, 'follow' contributors, 'comment' on and 'like' content, post your own content, and set up a tailored email digest for updates.