Marina Rodnina: “It is a great time to enter science.”

What’s currently your favourite molecule?
My favourite molecule is the ribosome, the molecular factory that synthesizes proteins in all living cells. The ribosome can be used to study a number of fundamental questions in biology, such as recognition, catalysis, movement, signalling, quality control or orchestrating complex networks of auxiliary factors and networks. In contrast to many other molecular complexes, the active sites of the ribosome consist of RNA, which significantly reduces the repertoire of the active groups that are involved in all these diverse functions. Because the ribosome is essential, evolution of the ribosome presents a relic from the RNA world that allows us to trace ancient biological history that is long eroded anywhere else. Finally, ribosome and protein synthesis – and their defects – play an important role in human diseases, which makes translation an important target in the search for novel drugs and therapies.
What are some of the challenges in your field right now?
One of the most challenging questions in our field right now is to understand the interplay between the pace of translation and the quality of protein folding on the ribosome during ongoing translation. It is known that elements in the mRNA, such as synonymous codon usage, sequence contexts and structural elements, can affect the codon-specific rate of translation. It is also known that changing the pace of translation can alter the way a nascent protein folds. However, so far the picture is complex: in some cases slower translation facilitates correct protein folding, while in the others it may lead to misfolding and protein aggregation. Some synonymous codons have a large effect while some have none at all. Moreover, nascent proteins remain very dynamic while attached to the ribosome and may fluctuate between different conformations. It is not known how these fluctuations or structural rearrangements affect the synthetic activity of the ribosome, although it is clear that there is a two-way communication between nascent-protein folding and ribosome activity. All these findings lead to the concept of a folding code, which entails information encoded in the mRNA beyond the meaning of codons in the genetic code. Deciphering this folding code is a major challenge for us and many other groups in the field.
Can group leaders still find time for hands-on research?
In my opinion, it is essential for any scientist to do as much hands-on research as possible. True discoveries are usually unexpected and need a trained eye that sees an exciting finding where everybody may conclude that the experiment did not work. To see a gold vein in a pile of unexplainable “rubbish”, one has to be intimately familiar with the way the experiments are done – best would be to do the experiments oneself. Also, the development of new techniques and troubleshooting is much more productive if the group leader is involved in the hands-on research. Finally, most young researchers entering science love their hands-on work (unlike administration, which is mostly frustrating) and have been selected as group leaders because they are good at doing science. So why not do things that you like and are good at? If there is a new bold idea, who should be better at testing it than the most experienced person in the lab, who is often the group leader. The time will come when committees take most of one’s time, so why not enjoy the time for doing research as long as possible.
What do you see as the main changes in the way researchers work in the past 20 years?
We now have fantastic research tools that one could only dream of 20 years ago. There is so much information from different “omics” databases that an emerging hypothesis can often be tested by just looking at existing data. Structural work has become much more accessible due to the progress of cryo-electron microscopy and the Alphafold predictions. Different imaging techniques provide unprecedented views of processes in living cells. All these versatile methods make it possible to ask challenging questions of outstanding biological importance and design experiments that provide best answers, rather than rely on a few feasible methods. It is a great time to enter science.
What worries you about life sciences research currently?
I am worried about two tendencies: the erosion of the peer review system and the excessive reliance of the scientific evaluation process on publication in very few top journals. There is nothing wrong with the peer review system as such. However, the way it works now, with essentially unlimited control of peer reviewers over the fate of a manuscript, is exaggerated and potentially hindering innovation and truly original work. None of the experiments to curb or limit this control worked very well so far. In my opinion, this problem needs some radical solutions very soon.
Introduction to Marina Rodnina’s work
Research summary
Research work in the Rodnina research group spans ribosome function and dynamics, regulation and fidelity of translation, and ribosome-catalyzed reactions. Current projects include cotranslational protein folding on the ribosome and inside of the membrane, recoding of the genetic information (such as frameshifting, stop-codon readthrough and translational bypassing) and the mechanisms of translation in human cells.
Lab webpage: https://www.mpinat.mpg.de/rodnina
Two recent/key papers:
Agirrezabala, X. et al. (2022) A switch from α-helical to β-strand conformation during co-translational protein folding. EMBO J. 41, e109175. https://doi.org/10.15252/embj.2021109175
Holtkamp, W. et al. (2015) Co-translational protein folding on the ribosome monitored in real time. Science 350,1104–1107. https://doi.org/10.1126/science.aad0344
More information on the FEBS Special Plenary Lecture at the 47th FEBS Congress
Marina Rodnina will deliver the FEBS Special Plenary Lecture at the 47th FEBS Congress on Tuesday 11th July 2022 on ‘The ribosome and the protein folding code of translation’: https://2023.febscongress.org/
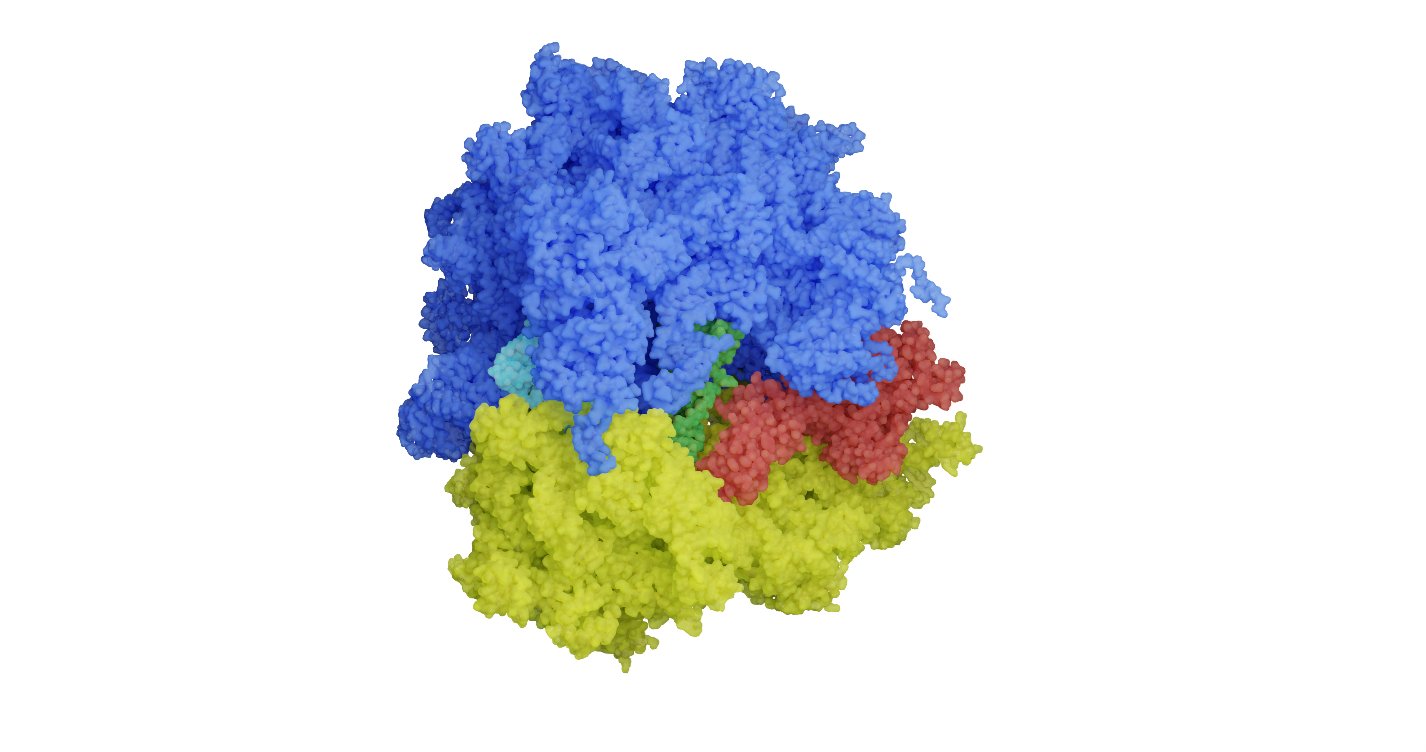
Top image of post: The structure of a bacterial ribosome, a molecular machine that synthesizes proteins in all cells and guides nascent proteins towards their correct cellular structure. Yellow and blue, the ribosomal subunits; green, tRNA; red, elongation factor G bound to the ribosome. Image from Marina Rodnina.





Join the FEBS Network today
Joining the FEBS Network’s molecular life sciences community enables you to access special content on the site, present your profile, 'follow' contributors, 'comment' on and 'like' content, post your own content, and set up a tailored email digest for updates.