How the Netherlands eScience Center contributes to cutting edge research in the Life Sciences

This post was co-written by Michiel Punt (Section Head of Life Sciences) and Giulia Crocioni (Research Software Engineer) at the Netherlands eScience Center.
The Netherlands eScience Center (eSC) is the national center that enables researchers from all domains to carry out cutting edge research through innovative research software. eSC was created in 2012 by NWO (the Dutch Research Council) and SURF (the organization for IT in Dutch education and research), following a recommendation by the Dutch government to establish a national organization for the innovation of ICT infrastructures and digital methodologies. eSC supports researchers from all disciplines in the use of advanced software, computing and digital technologies. It does this in two ways: by collaboratively designing sustainable software, and by building digital skills and expertise. eSC works with researchers based at Dutch Universities and research centres on a wide range of projects, helping to keep the Netherlands at the forefront of research.
In the past 20 years software has become indispensable in order to perform research. Software has become more advanced and requires more skills to utilize its full potential. At the eSC we collaborate with academic researchers to build powerful software and artificial intelligence, thereby enabling researchers to go beyond their own capabilities. We advocate for open science, aim to make software reusable, and are keen to deliver high quality code.
How do we organize ourselves?
We have research software engineers (RSE), who often have a background in computer science, physics, biology or a closely related discipline. Moreover, they all have experience in the academic field, often having obtained a PhD and several years of post-doctoral experience. A successful collaboration requires good communication right from the start of a project. Some domain knowledge helps to better understand each other and the research topic. We therefore have aligned ourselves along research disciplines, see figure 1. This way we ensure that the RSE’s working on a domain specific project understand the domain and the vocabulary.
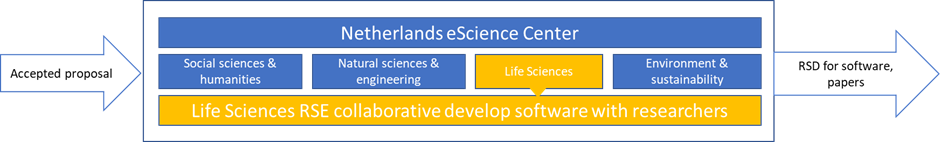
How can researchers collaborate with eSC?
The eSC offers various in-kind grants through Call for proposals. Researchers employed by Dutch knowledge institutes are eligible to submit a proposal. Each type of grant pursues a different goal. For instance, the Sustainable Software Call supports communities of researchers who require their software to meet higher quality standards to ensure the continuity and advancement of their research in the longer term. Another example, the Open eScience Call, contributes to state-of-the-art and innovative research that requires the development and application of advanced research software. Next to our Calls we often collaborate with researchers at a European level by joining international research consortiums. If you want to find out more, please contact the section head of your domain.
An example of a Life Sciences project
We would like to highlight a project which nicely represents the way we collaborate with researchers. The project “Personalized cancer vaccine design through 3D modelling boosted geometric learning” was accepted as an Open eScience Call project in late 2021. The project will last for 3 years and during that time the eSC offers 2 person-years of in-kind contribution. The eSC assigns the project to a team of 4 to 7 research software engineers. The assignment of a team is mainly based on experience with the project requirements, such as the software type or the data analysis to develop, and the affinity with the domain.
The broader scope of the project is to combine physics-based 3D modeling and data-driven deep learning for drug design and protein engineering. The team has already developed DeepRank and DeepRank-GNN, two deep learning frameworks for mining Protein-Protein Interactions (PPIs) through Convolutional Neural Networks (CNNs) and Graph Neural Networks (GNNs), respectively. In the current project, the team is developing DeepRank-Core, a package which unifies the above-mentioned softwares. DeepRank-Core is already being used to identify the most suitable mutated tumor peptides as candidates for personalized cancer vaccines. Preliminary experiments have yielded competitive results compared to state-of-the-art methods, and the framework presents significant advantages in terms of speed, storage requirements, and ease of use. The package is configured as flexible, easy-to-use, and open-source, it follows the community-approved FAIR principles for research software, and extensive documentation is already available. DeepRank-Core represents the result of a constant exchange of information between the eScience team engineers – the experts in software engineering and development’s best practices – and the research partners – the domain experts.
Research Software Directory
The Netherlands eScience center is a strong advocate for open science. This means that the software developed in collaboration with the eScience center should be accessible and reusable. To support this, together with partners we have recently set up the research software directory (RSD). The RSD is connected with GitHub and Zenodo, presents software together with associated data, research papers and researchers. Thereby the RSD helps to judge quickly the relevance of the software for you, encourages researchers to make software findable and accessible, and helps institutes to showcase developed software.
If you would like to find out more and/or are interested in setting up a (European) research consortium, please reach out to us.
Top photo provided by the Netherlands eScience Center





Join the FEBS Network today
Joining the FEBS Network’s molecular life sciences community enables you to access special content on the site, present your profile, 'follow' contributors, 'comment' on and 'like' content, post your own content, and set up a tailored email digest for updates.