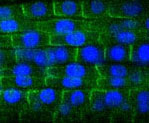
January - Issue 1
Pupylation of PafA or Pup inhibits components of the Pup-Proteasome System
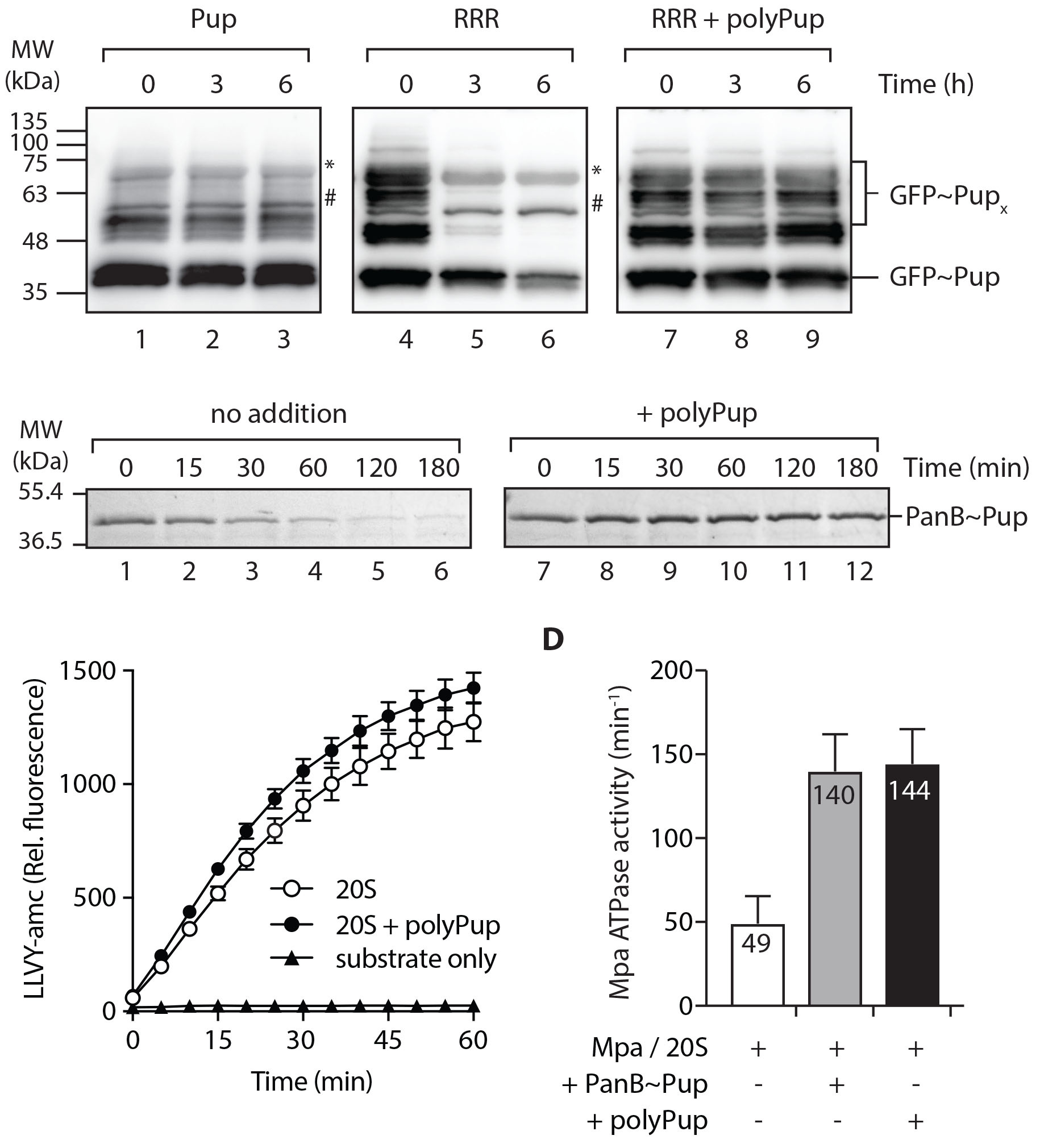
Dougan D.A and colleagues, FEBS Letters
The prokaryotic ubiquitin-like protein (Pup) is a functional analog of ubiquitin in Actinobacteria, and pupylation of proteins via the Pup ligase PafA leads to protein degradation via the bacterial 20S proteasome. Here, free or protein-linked polyPup chains primarily formed through K61 are shown to inhibit proteasomal function. Moreover, autopupylation of PafA in the absence of protein substrates inhibits ligase activity. The authors speculate that this negative feedback loop reversibly controls Pup availability and protein pupylation.
Kim J.H. and colleagues, FEBS Letters
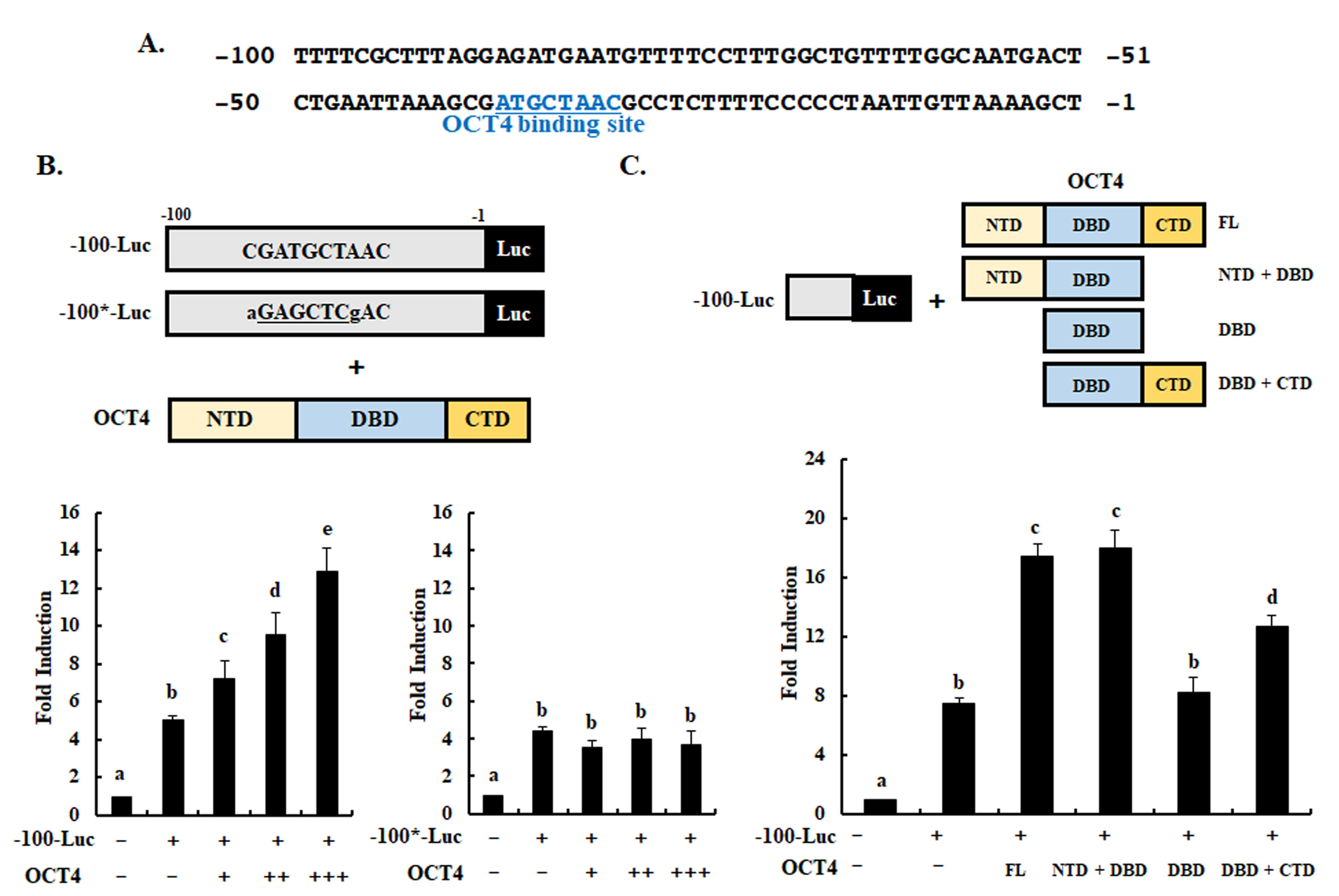
Highly malignant embryonal carcinoma (EC) cells maintain self-renewal and differentiation potential and share several signaling pathways with embryonic stem (ES) cells. In this study, Kim J.H. and colleagues show that the Cripto-1 gene, specifically over-expressed in EC and ES cells, is regulated by the transcription factor OCT4 in human EC NCCIT cells. To induce Cripto-1 expression OCT4 was shown to bind specifically to a novel OCT4-binding site within the Cripto-1 minimal promoter region.
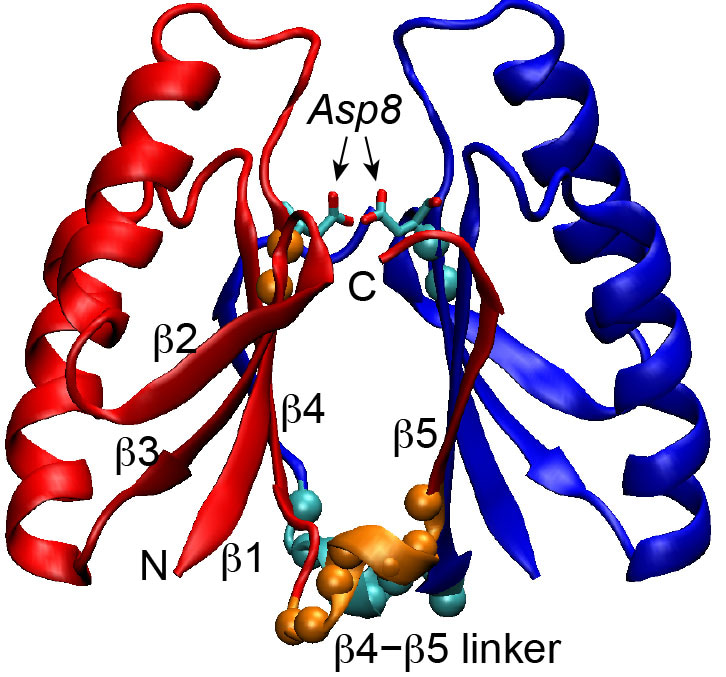
Suh J.Y. and colleagues, FEBS Letters
Cas2 of the CRISPR-Cas system functions as a scaffold protein and controls the length of the integrated spacer sequences. Beside this scaffold function, Cas2 exhibits nuclease activity against DNA and RNA substrates in a divalent metal ion dependent manner. Here, Suh and colleagues provide structural evidence that Cas2 predominantly adopts an inactive conformation in which Cas2 acts solely as a scaffold protein. However, significant conformational dynamics within the Cas2 active site allow for a conformational switch triggering its nuclease activity.
Doczi R. and colleagues, FEBS Letters
The localization of the auxin efflux carrier PIN1 at the plasma-membrane is needed to establish auxin gradients required during plant development. Here, the authors show that two environmentally stress-activated mitogen-activated protein kinases (MAPK), MPK4 and MPK6, phosphorylate conserved sites of PIN1, thereby reducing its plasma membrane localization. The authors propose an environmental adjustment of plant growth through the modulation of PIN1 plasma membrane localization by MAPKs.
January - issue 2
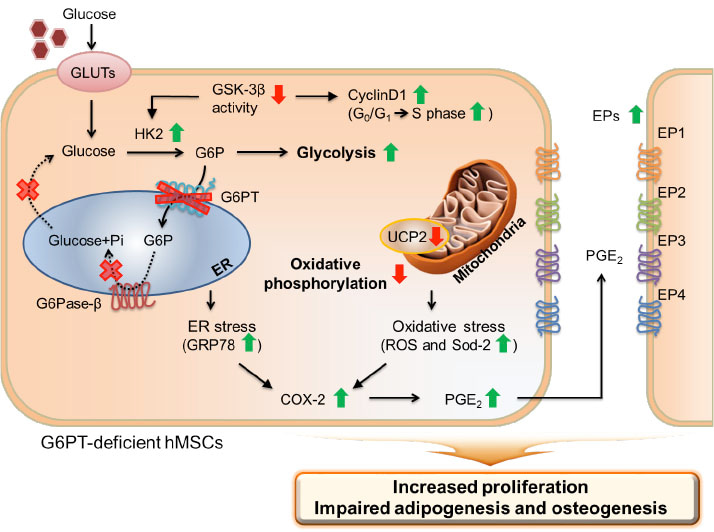
Jun, Hyun Sik and colleagues, FEBS Letters
The glucose-6-phosphate transporter (G6PT) is an endoplasmic reticulum (ER) membrane protein that shuttles cytoplasmic G6P into the ER lumen. Mutations in G6PT underlie glycogen storage disease type Ib, which is characterized by disrupted glucose homeostasis and immune deficiency. Here, G6PT is shown to regulate the proliferation and differentiation of human mesenchymal stem cells (hMSCs) via metabolic reprogramming and cyclooxygenase-2-mediated prostaglandin E2 secretion.
AID recruits the RNA exosome to degrade HIV-1 nascent transcripts through interaction with the
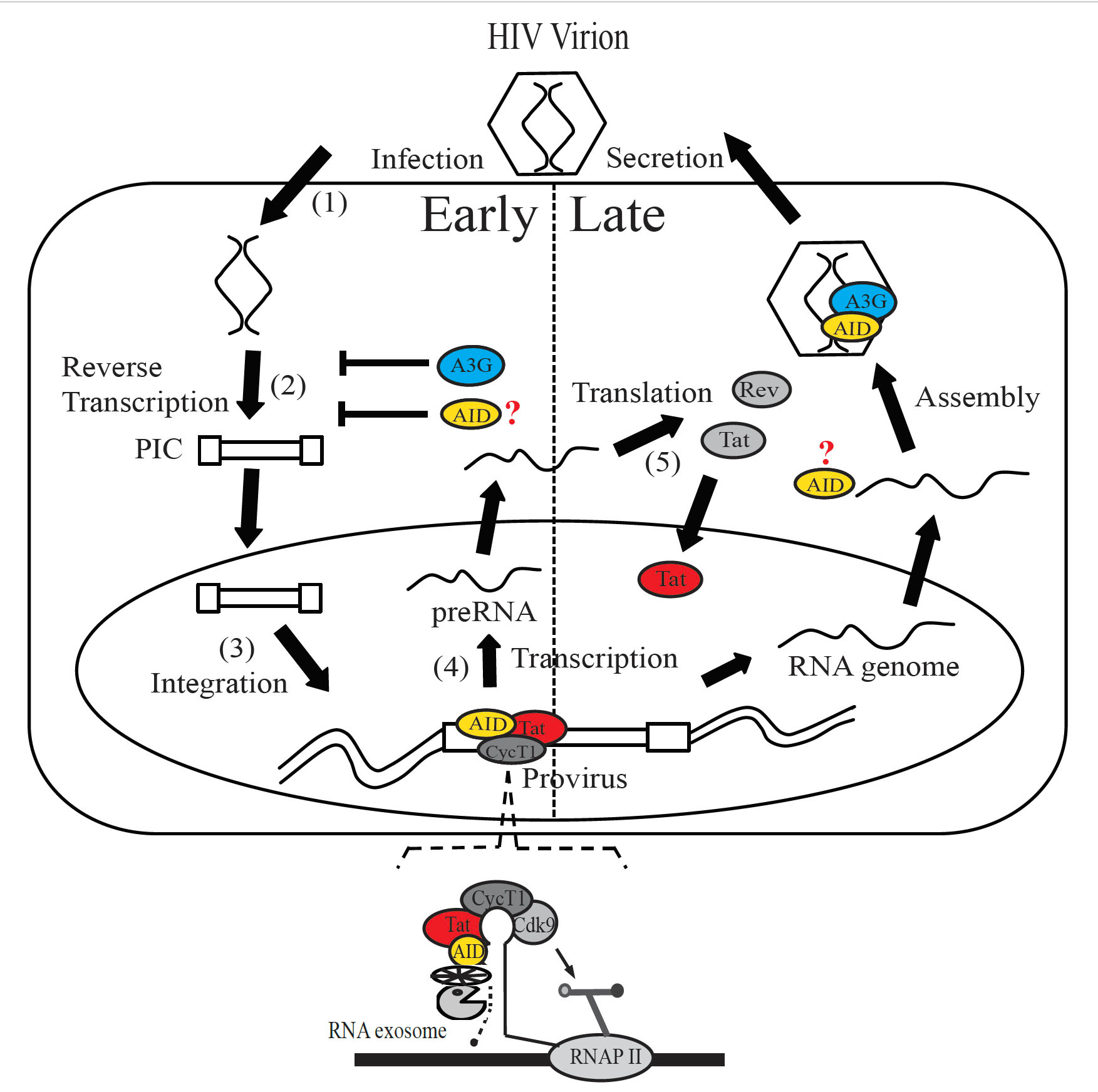
Shang, H. and colleagues, FEBS Letters
Activation-induced cytidine deaminase (AID) induces antibody diversification in B cells. Here, the authors reveal a role for AID in repressing HIV-1 transcription. AID is shown to interact with the HIV-1 Tat protein to mediate degradation of nascent HIV-1 transcripts by recruiting the RNA exosome. Intriguingly, the catalytically inactive mutant of AID had the same inhibitory effect, making it potentially suitable for use in gene therapy to suppress HIV-1 replication while avoiding wild-type AID deamination-mediated cellular genome instability.
Arabidopsis pollen extensins LRX are required for cell wall integrity during pollen tube growth
Muschietti, J. and colleagues, FEBS Letters
Polarized growth of pollen tubes involves constant remodelling of the cell wall. Here, Muschietti and colleagues show that the Arabidopsis leucine-rich repeat extensins (LRXs) 8-11, which are extracellular glycoproteins, function in maintaining cell wall integrity during pollen tube growth. Mutations in LRXs 8-11 result in abnormal pollen tubes and alterations in pollen germination and segregation ratio.




Join the FEBS Network today
Joining the FEBS Network’s molecular life sciences community enables you to access special content on the site, present your profile, 'follow' contributors, 'comment' on and 'like' content, post your own content, and set up a tailored email digest for updates.