Granular data: the AI era in cancer research
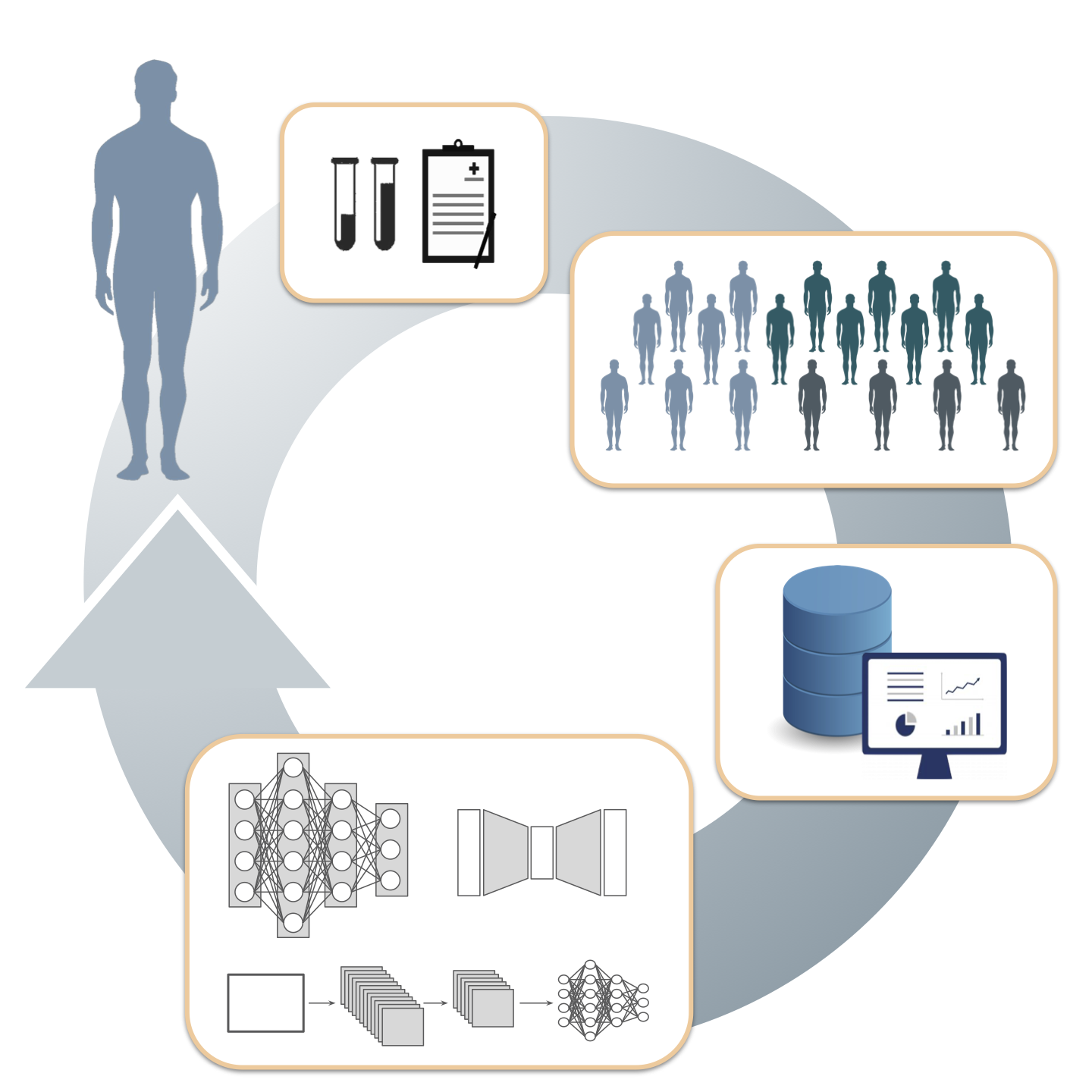
In their review article, Davide Cirillo, Iker Nuñez-Carpintero and Alfonso Valencia, address current challenges, limitations and solutions in converging big data in cancer research. Data granularity refers to the amount of detail observed in clinical and molecular data (e.g., demographic information, familial history, symptoms, comorbidities, histopathological features, immunohistochemistry, nucleic acids sequencing, biochemical analyses, digital images, experience measurements using digital devices) from patients with cancer. When dealing with granular data, Cirillo et al. emphasise on the need for implementing more standardised and systematic AI and ML approaches. The authors suggest that these approaches could be synergistically used to produce discriminative and generative models of the granular data continuum in cancer research.
Such granular datasets of combined biomarkers, DNA copy number aberrations (CNAs) and tumour heterogeneity of patients with colorectal liver metastasis were explored in the study of Kaja Berg and colleagues in order to improve prognostic patient stratification. The authors determined the mutational status of RAS, BRAFV600E and TP53, as well as the CNA profiles of liver lesions and tumours. By employing univariable and combined prognostic analyses on the molecular and clinical patient data, the authors demonstrate that CNA profiling can be used to predict the genomic and prognostic heterogeneity, as well as the clinical outcome of patients with colorectal liver metastasis carrying RAS, BRAFV600E and TP53 mutations.
A second research article in our current issue, linked big data analysis to prediction of therapeutic responses to PARP inhibitors (RAPRis) in patients with testicular germ cell tumors (TGCT). By using in silico tools, João Lobo et al. defined a specific set of genes acting in homologous recombination (HR) and pinpointed the CpG sites with negative correlation to HR gene expression. HR gene expression and promoter methylation was then correlated with PARPis treatment response and validated in vitro, concluding that HR gene promoter methylation may serve as a predictor of the therapeutic response to PARPis in patients with TGCT.
Next, the study of Leonie de Klerk and colleagues defined molecular predictive markers of responses to neoadjuvant chemoradiation in patients with localized oesophageal cancer. Pre-treatment biopsies were investigated by targeted next-generation DNA sequencing and promoter methylation analysis, and the resulting patterns were subsequently associated to histopathological responses and patient survival. By converging several granular datasets, the study identified candidate genetic biomarkers of response to neoadjuvant chemoradiotherapy (OAC, CSMD1, ETV4, SMURF1, SMARCA4, KRAS, GATA4, TP63, TFPI2, CDKN2A). These biomarkers could aid clinical decision-making for the treatment of patients with localized oesophageal cancer.
The ability to to predict therapeutic responses is key for stratifying patients eligible for immunotherapies, including therapies with immune-checkpoint inhibitors (ICIs). The central role of big data analysis for ICI responder stratification is showcased in the article by Joan Frigola et al. The authors used integrative analysis of genomic and transcriptomic features of long-term responders to ICIs. Whole-exome sequencing was used to estimate the tumor mutational burden (TMB) and the somatic CNAs (SCNAs) in patients with advanced non-small cell lung cancer (NSCLC), and these data were integrated with gene expression analysis. Long-term benefit following ICIs strongly associated to TMB. In addition, TMB, SCNA burden and PD-L1 expression were identified as complementary determinants of response to ICIs in patients with NSCLC.
Single cell transcriptomics have become indispensable in studying intra-tumoral heterogeneity and the tumour microenvironment (TME), thus providing an additional level of data granularity which can be employed in predicting the response to immunotherapy in patients with cancer. In their study, Jingtao Chen and colleagues have characterised 11,866 single T-cells from tumours or adjacent normal tissue of patients with oral squamous cell carcinoma (OSCC) by scRNAseq. Clustering analysis revealed 14 tumour-specific T-cell subpopulations of which exhausted CD8+ T-cells and regulatory CD4+ T-cells were enriched within the TME. Aberrant TOX expression regulated by transcription factor PRDM1, was outlined as a key molecular regulator of T cell dysfunction in OSCC tumours.




Join the FEBS Network today
Joining the FEBS Network’s molecular life sciences community enables you to access special content on the site, present your profile, 'follow' contributors, 'comment' on and 'like' content, post your own content, and set up a tailored email digest for updates.